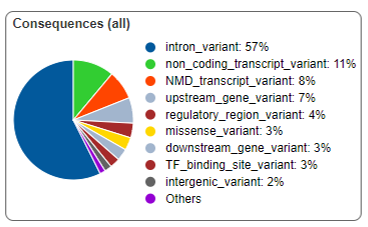
Exome sequencing is a widely used and cost-effective targeted sequencing method.
The application is used to enrich, sequence, and analyze the coding and regulatory regions of the genome, allowing the detection of disease-causing candidate variants (including SNPs, insertions and deletions), population genetics, and more.
For orders please read the samples delivery instructions, fill and send us the electronic sample sheet.
For additional information, please contact:
Liat Linde, tel 073-3785452 or 077-8875168
Rappaport building 073-3785221
Emerson building 077-8871387