The 10X Genomics Chromium Single Cell Gene Expression Solution provides a comprehensive, scalable solution for cell characterization and gene expression profiling of hundreds to millions of cells. The sequencing libraries are generated with the Chromium controller instrument and reagents from 10X Genomics, followed by sequencing on our Illumina sequencers.
The Chromium Single Cell Gene Expression Solution includes the GemCodetm Technology, which combines microfluidics with molecular barcoding, and custom bioinformatics software to enable 3’ mRNA counting from thousands of single cells.

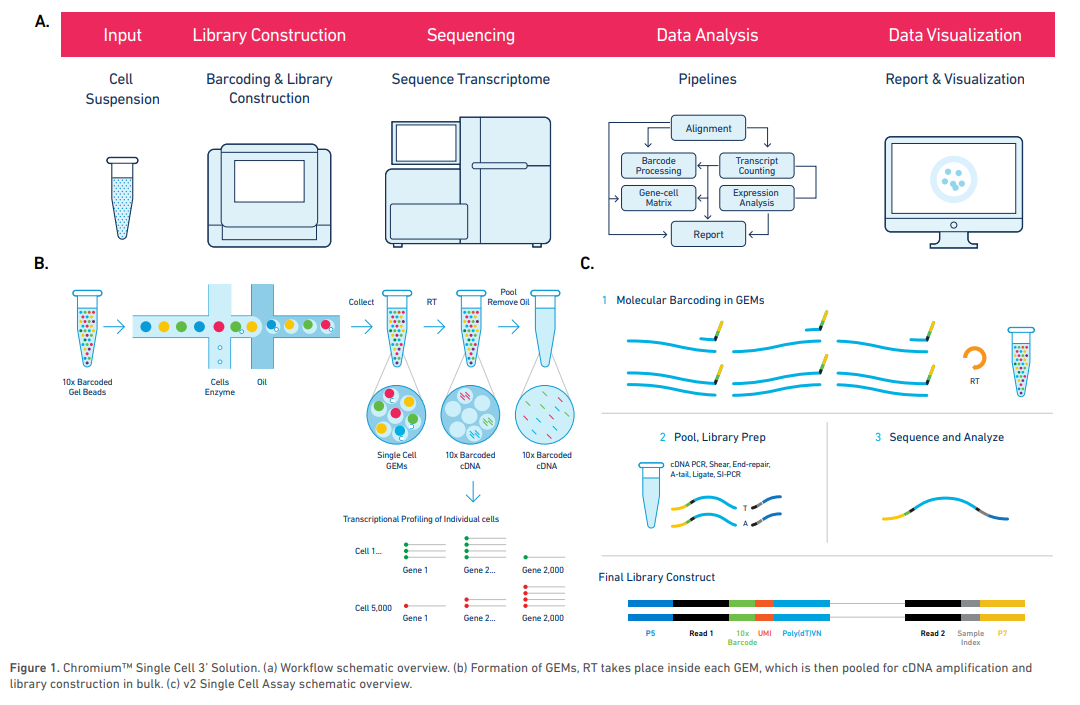
The 10X Genomics Chromium Single Cell 3’ Solution in brief
Single cells (or nuclei), reagents and a single Gel Bead containing barcoded oligonucleotides are encapsulated into nanoliter-sized GEMs (Gel Bead in emulsion) using the GemCodetm Technology. Lysis and barcoded reverse transcription of polyadenylated mRNA from single cells are performed inside each GEM. High-quality next generation sequencing libraries are finished in single bulk reaction and are compatible with Illumina sequencers. Finally, the Chromium™ Software Suite (Cell Ranger 2.0 and Loupe Cell Browser) is utilized for processing, analysis and visualization of single cell gene expression data.
10X Chromium Single Cell Features:
- Fast workflow from cell suspension to 3′-cDNA library.
- Captures 100-80,000+ cells (from up to 8 samples) in < 7 minutes.
- In addition to cell suspension samples also nuclei suspensions can be studied, enabling the analyses of brain tissues.
- Recovers up to ~65% of cells (typically 50%).
- Low doublet rate (~0.9% per 1,000 cells).
- Compatible with Illumina sequencers.
- For most applications an average of 50,000 reads per cell should be sequenced (for cell types with complex transcriptomes). For isolated nuclei 25,000 reads each are recommended.
- It is possible to work with cryo-preserved cells and methanol fixed, enabling safe sample shipping and batching.
- The cell size limit is comparatively high. Cells can have a diameter of up to 50 µm.
Samples submission requirements
The TGC team will prepare the single-cell RNA-seq libraries from cell suspension samples submitted by the research labs. The success of the library preparation is highly dependent on the state and quantification of the cells. Since the samples should be processed as quickly as possible, the experiments need to be planned and scheduled together with our staff.
